A new study in Open Biology is the first to demonstrate YAP dynamics in embryos using live imaging. We spoke to author Dr Bin Gu about what this means for embryonic research and the interdisciplinary collaboration behind it.
Please tell us about yourself and your/your group's research
I grew up and got my Ph.D. in Cell Biology in China, studying Embryonic Stem Cells. Then I went on to study mouse genetics and early embryo development with Dr. Janet Rossant as a postdoc in SickKids Hospital in Toronto, Canada. Working with my colleague Dr. Eszter Posfai and with support from Dr. Rossant, we found out that injecting CRISPR-Cas9 reagents into the 2-cell stage mouse embryos can strongly increase the efficiency of large fragment knock-in in mouse. We developed the 2-cell(2C)-Homologous Recombination (HR) – CRISPR technology, which allows routine generation knock-in reporter mouse models for live imaging. This is the pretext of the Open Biology article and probably the most important foundation of my whole independent research program now, as I have moved to the Institute of Quantitative Health Science and Engineering(IQ) at Michigan State University and started my independent Lab in 2020.
What is your article about?
The article is about a project I started in the Rossant Lab and carried on after I became independent. The project is a success because of an interdisciplinary collaboration between Rossant Lab and Hopyan Lab in SickKids Hospital, Sun Lab in the University of Toronto and my Lab. YAP protein (also known as Yes-associated protein 1) is one of the most important signaling proteins connecting biochemical and mechanical sensing processes. It transmits signals by repositioning its nuclear-cytoplasmic localization, when it is in nuclear, it activates gene transcription and is considered active. Because almost all previous research on YAP functions had relied on static analysis of fixed samples, it is an open question of how YAP behaves dynamically in an embryo and the functional significance of that behaviour. In this article, we described the first knock-in fusion reporter mouse line of Yap (Yap-emiRFP670) and revealed some unexpected dynamic behaviors of YAP protein in both pre- and post-implantation mouse embryos by live imaging. I will discuss these dynamic behaviors in the next question, but they point to interesting new avenues for studying YAP function in embryos. And I do hope this mouse model will be adopted and used by many scientists to reveal novel dynamic functions of YAP in the myriads of developmental, physiological, and pathological processes that YAP regulates.
What are the main points readers should take from the article?
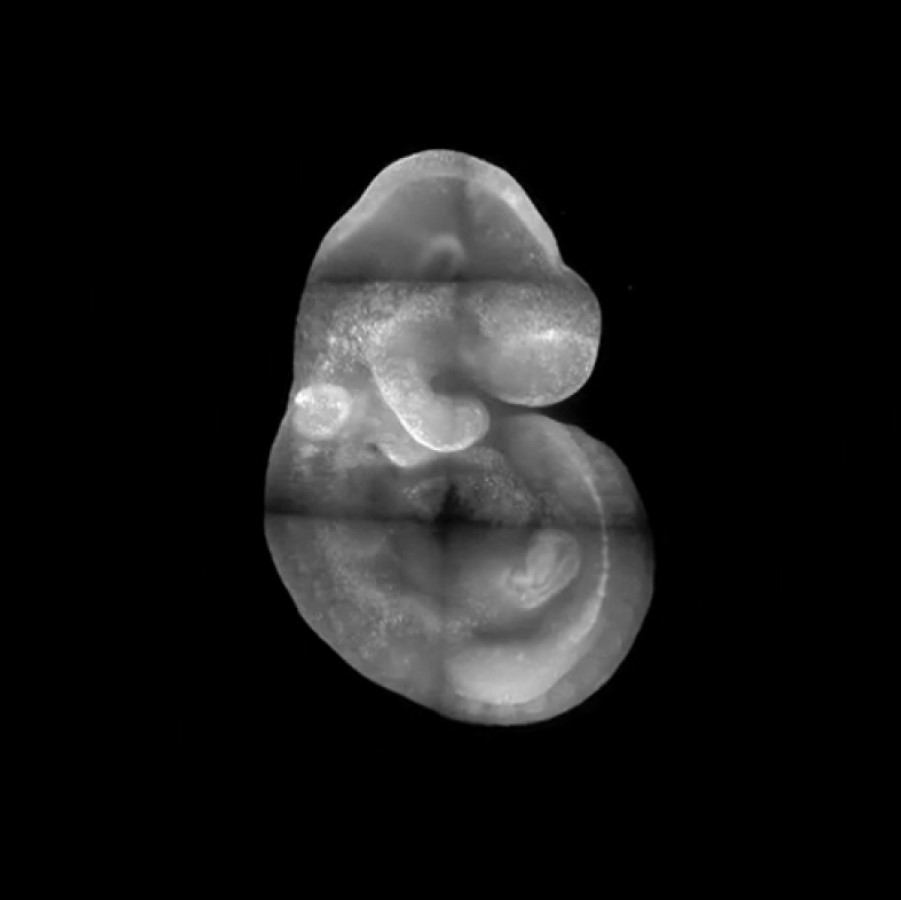
Perhaps the primary message the readers should take is that it is possible to live imaging YAP in vivo now, and lots of cool experiments can be done with this new tool. Aside from that, there are a few specific points. First, we spent a lot of effort in figuring out the fusion design that maintains the normal function of YAP protein in a knock-in mouse, and please take that into consideration when designing other fusion tagging in YAP. Second, we observed an intriguing mitotic reset behaviour of YAP protein during the first lineage segregation phase of mouse preimplantation embryos, meaning that YAP protein will always assume a nuclear localization after a cell division. The subcellular localization of YAP will then be adjusted based on the immediate environment of a blastomere in the later phase of the cell cycle. This behaviour can have some interesting implications on the cell cycle control of development and is worth further pursuing. Third, we presented direct observation that a group of sporadic cells with a strong nuclear YAP signal conducts persistent cell migration within the head region of early organogenesis (E8.0 to 8.5) stage embryos. This observation indicates a role of YAP activity in directing long-range cell migration. The lineage identity of those cells and the causal relationship between YAP activity and the persistent cell migration in these cells merit further investigation.
Did anything surprise you while conducting your study?
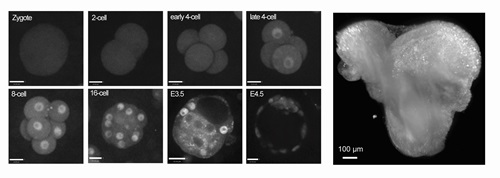
There are always many surprises when you see any developmental process unfold in life for the first time. Probably the most surprising is the mitotic reset behaviour of YAP in preimplantation embryos, and the lack of correlation between nuclear YAP and the expression of the trophectoderm marker Cdx2 at early 16-cell stage we discussed in the Open Up section in the article. These observations contradict some former models of first lineage segregation in mouse embryos and merit further investigation.
Your manuscript was invited by our Preprint Editorial Team, what was your experience publishing via this route?
It is absolutely a great and enjoyable experience to publish via the Preprint Editorial Team invitation. I am thankful to the preprint editors and highly recommend anyone to consider this route if they are contacted.
What's next for you and your research?
Overall, I like to build mouse models to address previously challenging or impossible problems. The Lab is recently working on new chromosome translocation technology for modeling fusion oncogenes, and further engineering the Auxin Inducible Degradation Mouse Model we recently described in a preprint to manipulate endogenous genes such as YAP/TAZ in living mice with high lineage and temporal precision. I hope that with these new technologies, we will be able to make discoveries that will open up new avenues in research distinctively from well-established models, similar to the YAP reporter in the present article.
Do you have an exciting new discovery that you would like to publish in Open Biology? Find out more about our author benefits and submission process.
---------------------------------------------------------------Movie: Time lapse movie revealed a population of cells with strong nuclear YAP signal migrating within the head region of mouse embryos. Credit Bin Gu
Figure: The dynamic behaviour of cells with active YAP signalling (still from movie).




